This web page was produced as an assignment for Genetics 564 at UW-Madison in Spring 2014.
What is Protein Homology?
Two proteins highly similar due to common ancestry are classified as homologous to one another. These can help us understand evolutionary events at a protein level, as well as give us tools to study disease. When a homolog is identified in a model organism with highly conserved function and sequence, scientists can use this to better understand protein functions, even if genomic DNA sequences are not highly conserved.
PEX1 Protein Sequences
|
Bos taurus (Cow, domestic)
Peroxisome biogenesis factor 1 (PEX1) Accession: NP_001179000.1 Length: 1,281 aa Danio rerio (Zebrafish) Peroxisome biogenesis factor 1 (PEX1) Accession: NP_001164306.1 Length: 1,237 aa Arabidopsis thaliana (Thale Cress) Peroxisome biogenesis protein 1 (PEX1) Accession: NP_196464.2 Length: 1,130 aa |
Rattus norvegicus (Brown Rat)
Peroxisome biogenesis factor 1 (PEX1) Accession: NP_001102690.1 Length: 1,283 aa Drosophila melanogaster (Fruit Fly) Peroxin 1 (PEX1) Accession: NP_652016.1 Length: 1,006 aa Saccharomyces cerevisiae (Yeast) AAA family ATPase peroxin 1 Accession: NP_012724.1 Length: 1,043 aa |
Mus musculus (House Mouse)
Peroxisome biogenesis factor 1 (PEX1) Accession: NP_082053.1 Length: 1,244 aa Caenorhabditis elegans (Roundworm) Peroxin (PRX-1) Accession: BAB62002 Length: 996 aa |
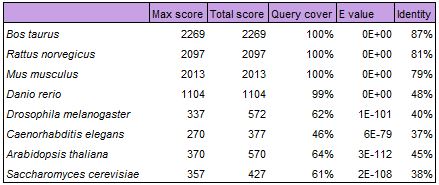
BLAST Alignment
The following table shows percent identity data from a protein BLAST alignment using blastp [1]. Click here to download the alignment.
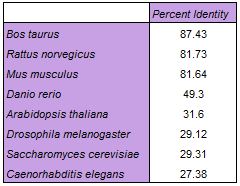
Clustal Omega Alignment
The table below shows percent identity data extrapolated from a percent identity matrix (download) from Clustal Omega. A percent identity matrix assigns identity values between each organism investigated [1]. Click here to download the alignment.
T-Coffee Alignment
Click here to download the alignment from T-Coffee. In this file type, colors denote the accuracy of the alignment [2,3].
Discussion
The alignment of these protein sequences using the various methods described has shown a high amount of conservation among mammals, and a moderate amount of conservation in other non-mammals listed with a fairly high level of certainty. This is true across three different alignment methods. Homology appears to decrease drastically in invertebrates.
[1] Sievers F, Wilm A, Dineen D, Gibson T, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, Thompson J, Higgins D, Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega, MSB 7 (2011) 1-6. PMID: 21988835
[2] Di Tommaso P, Moretti S, Xenarios I, Orobitg M, Montanyola A, Chang JM, Taly JF, Notredame C, T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension, Nucleic Acids Res. 39 (2011) W13-7. PMID:21558174
[3] Notredame C, Higgins D, Heringa J, T-Coffee: A novel method for fast and accurate multiple sequence alignments. JMB
302 (2000) 205-217. PMID:10964570
[2] Di Tommaso P, Moretti S, Xenarios I, Orobitg M, Montanyola A, Chang JM, Taly JF, Notredame C, T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension, Nucleic Acids Res. 39 (2011) W13-7. PMID:21558174
[3] Notredame C, Higgins D, Heringa J, T-Coffee: A novel method for fast and accurate multiple sequence alignments. JMB
302 (2000) 205-217. PMID:10964570