This web page was produced as an assignment for Genetics 564 at UW-Madison in Spring 2014.
What is Gene Homology?
Two genes highly similar due to common ancestry are classified as homologous to one another. These can help us understand evolutionary events at a genetic level, as well as give us tools to study disease. When a homolog is identified in a model organism with highly conserved function and sequence, scientists can use this to better understand gene and the disease mutations to it may cause.
PEX1 Genomic Sequences
The PEX1 gene is highly conserved across many organisms. Below are links to select model organism sequences of PEX1 homologs.
|
Bos taurus (Cow, domestic)
peroxisome biogenesis factor 1 (PEX1) Accession: AC_000161.1 Length: 81,806 bp Danio rerio (Zebrafish) peroxisome biogenesis factor 1 (PEX1) Accession: NC_007130.5 Length: 32,717 bp Arabidopsis thaliana (Thale Cress) peroxisome biogenesis protein 1 (PEX1) Accession: NC_003076.8 Length: 9,272 bp |
Rattus norvegicus (Brown Rat)
peroxisome biogenesis factor 1 (PEX1) Accession: NC_005103 Length: 50,374 bp Drosophila melanogaster (Fruit Fly) peroxin 1 (PEX1) Accession: NT_037436 Length: 5,502 bp Saccharomyces cerevisiae (Yeast) AAA family ATPase peroxin 1 Accession: NC_001143.9 Length: 4,072 |
Mus musculus (House Mouse)
peroxisome biogenesis factor 1 (PEX1) Accession: NC_000071.6 Length: 41,165 bp Caenorhabditis elegans (Roundworm) peroxin (PRX-1) Accession: NC_003284.8 Length: 16,418 |
BLAST Alignment
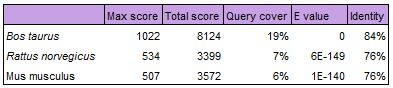
The table below displays results of a Nucleotide BLAST alignment of the human PEX1 gene. All of the organisms listed above were used, and Megablast was used to find highly similar sequences. Those not listed did not show significant alignment similarity [1].
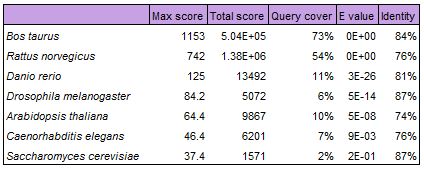
The following table display results of another Nucleotide BLAST alignment of the human PEX1 gene. The blastn algorithm was used instead to identify somewhat similar sequences.
Kalign Alignment
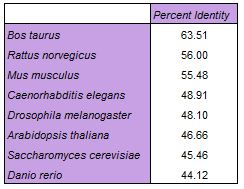
The Kalign sequence alignment tool was used since it is capable of handling larger sequences than others, such as Clustal Omega. The table below shows percent identity data extrapolated from a percent identity matrix [2]. The alignment can be downloaded here.
Discussion
It is difficult to ascertain any solid conclusions regarding the gene homology of PEX1, since its length makes it difficult for many alignment algorithms to handle. There is also a significant difference in gene sequence length across the set of organisms analyzed. However, it does appear that the PEX1 gene sequence is semi-conserved at least in the mammalian organisms analyzed. It should be noted that the PEX1 protein is more highly conserved.
[1] McGinnis S, Madden T, BLAST: at the core of a powerful and diverse set of sequence analysis tools, Nuc. Acids Res. 32 (2004) W20-W25. PMID: 15215342
[2] Lassmann T, Sonnhammer E, Kalign-an accurate and fast multiple sequence alignment algorithm, BMC Bioinf. 6 (2005) PMID: 16343337
[2] Lassmann T, Sonnhammer E, Kalign-an accurate and fast multiple sequence alignment algorithm, BMC Bioinf. 6 (2005) PMID: 16343337