This web page was produced as an assignment for Genetics 564 at UW-Madison in Spring 2014.
What is Gene Ontology (GO)?
Gene ontology addresses three areas of gene description:
- Biological process: describes what organismal, tissue, and/or molecular processes the gene product is involved in
- Cellular component: describes the location of the gene product within the cell or its surroundings
- Molecular function: describes the functions of the gene product [1]
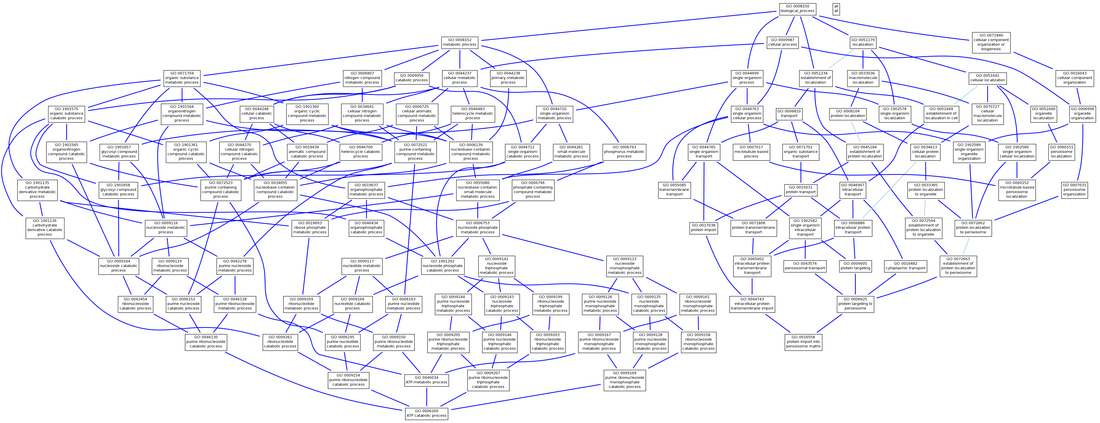
PEX1 Biological Process
Human PEX1 is implicated in six different biological processes:
- ATP catabolic process (GO:0006200), child of metabolic process (GO:0008152)
- microtubule-based peroxisome localization (GO:0060152)
- peroxisome organization (GO:0007031)
- protein import into peroxisome matrix (GO:0016558)
- protein targeting to peroxisome (GO:0006625)
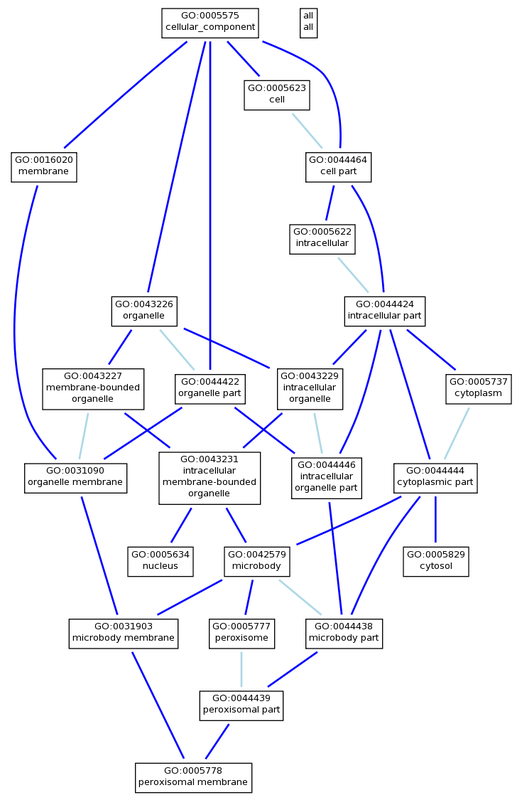
PEX1 Cellular Component
Human PEX1 is associated with seven different cellular components:
- cytoplasm (GO:0005737)
- cytosol (GO:0005829)
- intracellular membrane-bounded organelle (GO:0043231)
- NOT nucleolus (GO:0005730)
- nucleus (GO:0005634)
- peroxisomal membrane (GO:0005778)
- peroxisome (GO:0005777)
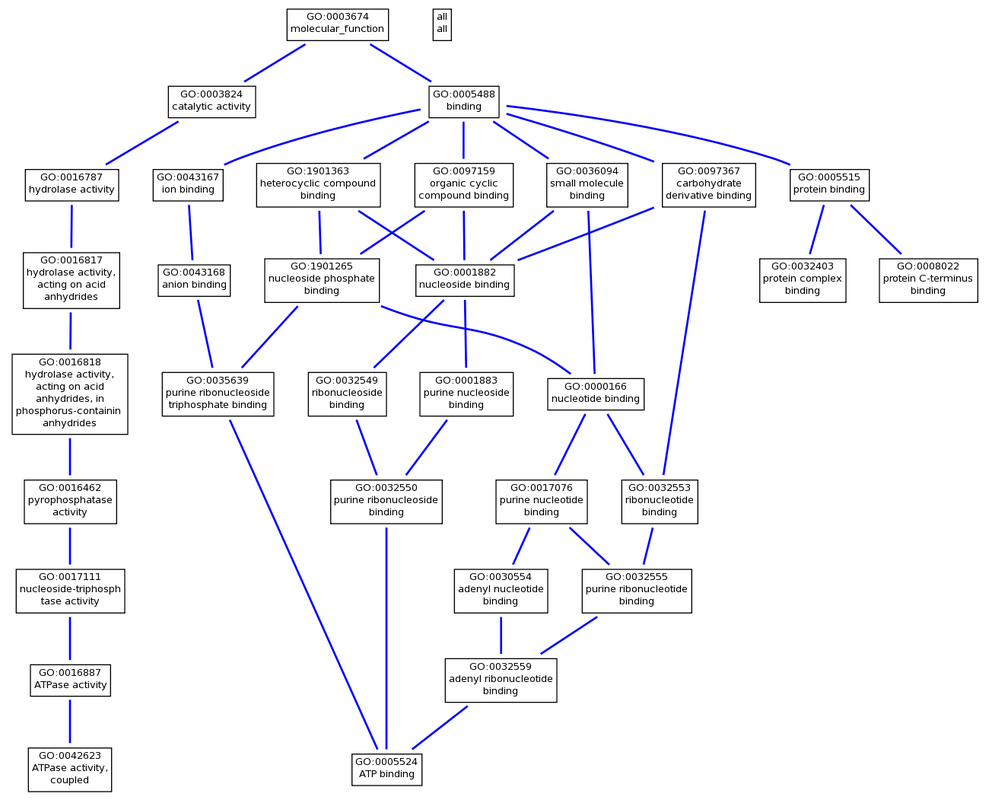
PEX1 Molecular Function
Human PEX1 is associated with five different molecular functions:
- ATP binding (GO:0005524)
- ATPase activity, coupled (GO:0042623)
- protein binding (GO:0005515)
- protein C-terminus binding (GO:0008022)
- protein complex binding (GO:0032403)
Discussion
The GO terms found using AmiGO for human PEX1 show a great deal of consistency with the primary literature, protein domain composition, and PEX1 mutation disease phenotypes. Since human PEX1 contains two AAA ATPase domains, it makes sense that the gene product would take part in ATP binding and ATPase activities as shown in the molecular function annotations. Since patients with mutant PEX1 have a buildup of waste products normally broken down by peroxisomes, it also makes sense that the biological process annotations include peroxisome organization, microtubule-based peroxisome localization, protein import into peroxisome matrix, and protein targeting to peroxisome, along with the cellular component of peroxisome. The GO trees generated elucidate the relationships between the annotations of PEX1.
[1] Hill D, Berardini T, Howe D, Van Auken K, Representing Ontogeny Through Ontology: A Developmental Biologists's Guide to the Gene Ontology, Mol Reprod Dev. 77 (2010) 314-329. PMID: 19921742