This web page was produced as an assignment for Genetics 564 at UW-Madison in Spring 2014.
What is a protein interaction network?
Protein interaction networks are graphical ways of representing protein-protein interactions. These interactions are deduced from experimental data and inferred from theoretical data, and can be direct or indirect interactions. These diagrams can give clues as to the function of the protein of interest. For example, if the protein interacts with geneX involved in processY, then that protein is probably involved in that process.
PEX1 protein interaction networks
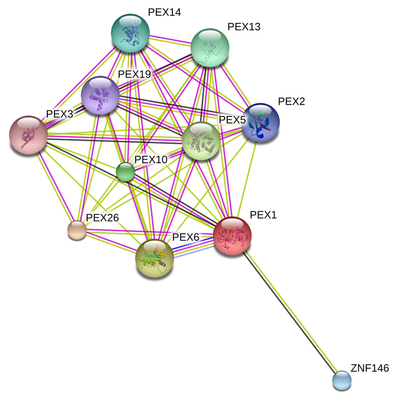
Below is an protein interaction network for PEX1 generated using STRING, and online database of protein interactions [1]. The interactions displayed are derived from different kinds of evidence such as experimental evidence, genomic context, and co-expression.
PEX1 is clearly interacting with most of the peroxin genes involved in peroxisomal biogenesis. This is consistent with previous research and proposed mechanisms of this process. One protein interactor, zinc finger protein 146 (ZNF146) was unexpected. ZNF146 is involved in the regulation of transcription via DNA and zinc ion binding. It also binds heparin. Basically, heparin is a glycosaminoglycan, a type of polysaccharide (sugar) that are often found of the extracellular matrix of the cell. Heparin inhibits blood coagulation by binding to antithrombin, a protease inhibitor that can then bind and inhibit thrombin, a protease necessary for proper clotting. However, it is unclear how ZNF146 is involved in this process [2]. It is also worth noting that ZNF146 localizes to the nucleus [3].
Discussion
The preceding interaction network further supports that PEX1 is involved in peroxisomal biogenesis, since it interacts with other proteins directly involved in this process. The unexpected protein interactor, ZNF146, is puzzling, especially when taking into account its localization to the nucleus. Since PEX1 localizes to the peroxisome, it could be that ZNF146 is somehow interacting with PEX1 while it is being transcribed or transported out of the nucleus, but this is purely speculation. This particular interaction is only supported by coexpression data, which just means that both PEX1 and ZNF146 are expressed together, so it will be interesting to see if both proteins exhibit a functional interaction with one another.
[1] Jensen et al. Nucleic Acids Res. 2009 37(Database issue): D412-6 <http://string-db.org/>.
[2] Nelson, David L., and Michael M. Cox. Principles of Biochemistry. New York: W.H. Freeman and Company, 2008. Print.
[3] Gene Ontology of ZNF146. Obtained from AmiGO 2 Database. <http://amigo.geneontology.org/amigo/gene_product/UniProtKB:Q15072>. Refers to: GO_REF:0000033, GO_REF:0000037, PMID:8665923, PMID:8665923, PMID:8665923. Last updated 5-11-2014.
[2] Nelson, David L., and Michael M. Cox. Principles of Biochemistry. New York: W.H. Freeman and Company, 2008. Print.
[3] Gene Ontology of ZNF146. Obtained from AmiGO 2 Database. <http://amigo.geneontology.org/amigo/gene_product/UniProtKB:Q15072>. Refers to: GO_REF:0000033, GO_REF:0000037, PMID:8665923, PMID:8665923, PMID:8665923. Last updated 5-11-2014.