This web page was produced as an assignment for Genetics 564 at UW-Madison in Spring 2014.
What is a protein domain?
A domain is a span of amino acids that serves as a structural and sometimes functional unit of a protein. These are used to gain insight into protein function as well as evolutionary conservation [1].
PEX1 Protein Domains
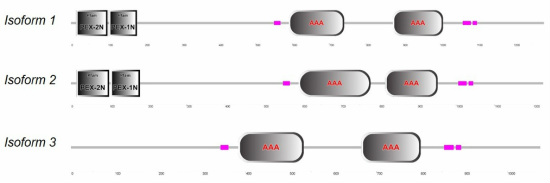
The PEX1 protein contains four domains: two peroxisomal biogenesis factor 1, N-terminal domains (PEX-2N and PEX-1N domains) and two ATPase Associated with diverse cellular Activities domains (AAA domains) [2,3].
PEX1 Isoform Differences
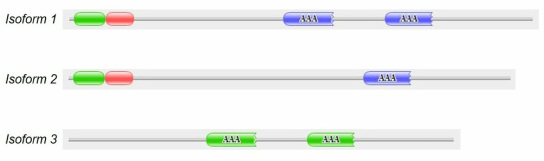
Human PEX1 has three different isoforms that arise from alternative splicing of mRNA. Their domain composition differs. Note that isoform 3 lacks both PEX-2N and PEX-1N [2,3].
PEX1 Domain Homolog Differences
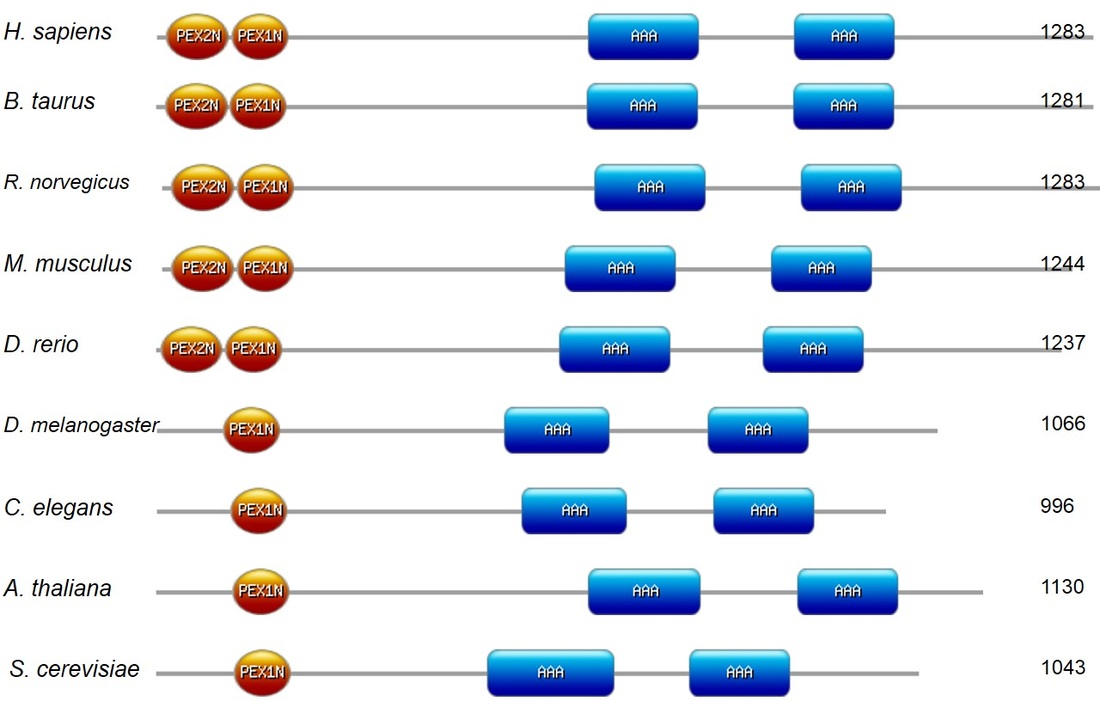
Using SMART and Pfam, protein domains in PEX1 homologs among select model organisms were identified. Shown below is a domain alignment generated using PROSITE's My Domain builder. Note that all nine organisms analyzed contain two AAA ATPase domains and one PEX-1N domain. All vertebrates analyzed contain a PEX-2N domain.
PEX-2N and PEX-1N Domains
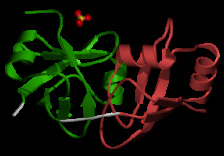 Figure 6: Mouse PEX1 N-terminal domains.
Figure 6: Mouse PEX1 N-terminal domains.
These domains take on double psi beta-barrel folds, and make up the N-terminus of PEX1. The PEX-2N (amino acids 14-99) and PEX-1N (amino acids 103-179) domains make up lobes (green and red, respectively, on the structure displayed) that flank a linker region. AAA ATPase proteins with similar N-terminal domains, Valosin-containing protein (VCP) and N-ethylmaleimide sensitive factor (NSF), contain hydrophobic residues in this linker region, allowing them to interact with ubiquitin. However, the linker region of PEX1 contains considerably fewer hydrophobic residues. It is hypothesized that this linker region still interacts with ubiquitin, but in a different way from VCP and NSF [4]. Ubiquitin is a small post-translational modifier protein that when bound, can mark the protein for degradation, affect its localization, activity, and/or interactions with other proteins. It also could be binding to phosphoinositides in peroxisomal membranes [5].
AAA ATPase Domains
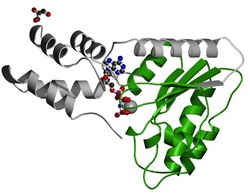 Figure 7: Chinese hamster NSF AAA ATPase domain.
Figure 7: Chinese hamster NSF AAA ATPase domain.
AAA protein domains contain two subdomains. The first is a P-loop NTPase at the N-terminus with an alpha-beta-alpha structure. Taking on a Rossman fold, this subdomain has motifs that make it possible for it to bind and hydrolyze ATP. The glycine rich Walker A motif (GXXXXFK(T/S)) interacts with phosphates of nucleotides and a magnesium ion. The Walker B motif ((R/K)XXXXGXXXXLhhhhD, where h's are hydrophobic residues) contains a span of negatively-charged amino acid residues, followed by hydrophobic ones. This motif is essential for magnesium ion interaction and ATP hydrolysis [6]. The second makes up a poorly-conserved C-terminus with an alpha structure [7].
PEX1 Motifs
Protein motifs are often used to predict protein function and can be used to identify related proteins.
Using MEME, three protein motifs were identified with conservation across all model organisms analyzed [8]. The following domain and motif images of human PEX1 were generated using PROSITE's My Domains builder. It's important to note that all three motifs identified lie within the AAA ATPase domains.
Using MEME, three protein motifs were identified with conservation across all model organisms analyzed [8]. The following domain and motif images of human PEX1 were generated using PROSITE's My Domains builder. It's important to note that all three motifs identified lie within the AAA ATPase domains.
Motif 1
Motif 1 is located within the second AAA ATPase domain in human PEX1.
Motif 2
Motif 2 is located towards the C-terminal end of the second AAA ATPase domain in human PEX1.
Motif 3
Motif 3 is located at the N-terminal ends of both AAA ATPase domains in human PEX1. Based on its location within the AAA ATPase domains and its amino acid composition, it is likely that positions 8-15 in this motif make up the Walker A motif described above.
Discussion
Protein domains and motifs can be used to infer a protein's function. PEX1 contains two N-terminal domains, two AAA ATPase domains, and three identified motifs, one of which is most likely a Walker A motif found in AAA ATPase domains.
The N-terminal domains have been shown to bind specific lipids in vitro [5]. This presumably affects the localization of PEX1. It is important to note that the PEX-2N domain is not conserved in D. melanogaster, C. elegans, A. thaliana, and S. cerevisiae. AAA ATPase domains have various functions in different proteins, but they carry these out using energy from ATP hydrolysis. These domains somehow contribute to the role PEX1 plays in peroxisome protein import by interactions with PEX5, PEX6, and PEX26 [9].
Literature states that PEX1 is anchored to the peroxisomal membrane via PEX26, however, Shiozawa et al., have shown that PEX1 also binds to lipids directly [5, 9]. It would be interesting to see if and how these two interactions cooperate to properly localize PEX1. Since the PEX-2N domain is not conserved across all of the species analyzed, it may not be necessary for proper lipid binding, and therefore, peroxisome localization in higher organisms.
The N-terminal domains have been shown to bind specific lipids in vitro [5]. This presumably affects the localization of PEX1. It is important to note that the PEX-2N domain is not conserved in D. melanogaster, C. elegans, A. thaliana, and S. cerevisiae. AAA ATPase domains have various functions in different proteins, but they carry these out using energy from ATP hydrolysis. These domains somehow contribute to the role PEX1 plays in peroxisome protein import by interactions with PEX5, PEX6, and PEX26 [9].
Literature states that PEX1 is anchored to the peroxisomal membrane via PEX26, however, Shiozawa et al., have shown that PEX1 also binds to lipids directly [5, 9]. It would be interesting to see if and how these two interactions cooperate to properly localize PEX1. Since the PEX-2N domain is not conserved across all of the species analyzed, it may not be necessary for proper lipid binding, and therefore, peroxisome localization in higher organisms.
[1] "What are protein domains?" EMBL-EBI. European Bioinformatics Institute, 2014. Web. 23 Feb. 2014. <http://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains>.
[2] Letunic I, Copley R, Pils B, Pinkert S, Schultz J, Bork P, SMART 5: domains in the context of genomes and networks, Nuc. Acid Res. 34 (2006) D257-260. PMID: 16381859
[3] Finn R, Tate J, Mistry J, Coggill P, Sammut S, Hotz H, Ceric G, Forslund K, Eddy S, Sonnhammer E, Bateman A, The Pfam protein families database, Nuc. Acids Res. 36 (2008) D281-288. PMID: 18039703
[4] Shiozawa K, Maita N, Tomii K, Seto A, Goda N, Akiyama Y, Shimizu T, Shirakawa M, Hiroaki H, Structure of the N-terminal domain of PEX1 AAA-ATPase. Characterization of a putative adaptor-binding domain, J. Biol. Chem. 279 (2004) 50060-50068. PMID 15328346
[5] Shiozawa K, Goda N, Shimizu T, Mizuguchi K, Kondo N, Shimozawa N, Shirakawa M, Hiroaki H, The common phospholipid-binding activity of the N-terminal domains of PEX1 and VCP/p97, FEBS J. 273 (2006) 4959-4971. PMID: 17018057
[6] Snider J, Thibault G, Houry W, The AAA+ superfamily of functionally diverse proteins, Genome Biol. 9 (2008) epub.
PMID: 18466635
[7] Neuwald A, Aravind L, Spouge J, Koonin E, AAA+: A class of chaperone-like ATPases associated with the assembly, operation, and disassembly of protein complexes, Genome Res. 9 (1999) 27-43. PMID: 9927482
[8] Bailey T, Elkan C, Fitting a mixture model by expectation maximization to discover motifs in biopolymers, Proc Int Conf Intell Syst Mol Biol. 2 (1994) 28-36. PMID: 7584402
[9] Hans R. Waterham, Merel S. Ebberink, Genetics and molecular basis of human peroxisome biogenesis disorders, Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease, Volume 1822, Issue 9, September 2012, Pages 1430-1441. PMID: 22871920.
Image References
secondary structure generated with: http://prosite.expasy.org/mydomains/
N-terminal secondary structure: http://pfam.xfam.org/structure/1WLF
AAA ATPase secondary structure: http://pfam.xfam.org/structure/1D2N
[2] Letunic I, Copley R, Pils B, Pinkert S, Schultz J, Bork P, SMART 5: domains in the context of genomes and networks, Nuc. Acid Res. 34 (2006) D257-260. PMID: 16381859
[3] Finn R, Tate J, Mistry J, Coggill P, Sammut S, Hotz H, Ceric G, Forslund K, Eddy S, Sonnhammer E, Bateman A, The Pfam protein families database, Nuc. Acids Res. 36 (2008) D281-288. PMID: 18039703
[4] Shiozawa K, Maita N, Tomii K, Seto A, Goda N, Akiyama Y, Shimizu T, Shirakawa M, Hiroaki H, Structure of the N-terminal domain of PEX1 AAA-ATPase. Characterization of a putative adaptor-binding domain, J. Biol. Chem. 279 (2004) 50060-50068. PMID 15328346
[5] Shiozawa K, Goda N, Shimizu T, Mizuguchi K, Kondo N, Shimozawa N, Shirakawa M, Hiroaki H, The common phospholipid-binding activity of the N-terminal domains of PEX1 and VCP/p97, FEBS J. 273 (2006) 4959-4971. PMID: 17018057
[6] Snider J, Thibault G, Houry W, The AAA+ superfamily of functionally diverse proteins, Genome Biol. 9 (2008) epub.
PMID: 18466635
[7] Neuwald A, Aravind L, Spouge J, Koonin E, AAA+: A class of chaperone-like ATPases associated with the assembly, operation, and disassembly of protein complexes, Genome Res. 9 (1999) 27-43. PMID: 9927482
[8] Bailey T, Elkan C, Fitting a mixture model by expectation maximization to discover motifs in biopolymers, Proc Int Conf Intell Syst Mol Biol. 2 (1994) 28-36. PMID: 7584402
[9] Hans R. Waterham, Merel S. Ebberink, Genetics and molecular basis of human peroxisome biogenesis disorders, Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease, Volume 1822, Issue 9, September 2012, Pages 1430-1441. PMID: 22871920.
Image References
secondary structure generated with: http://prosite.expasy.org/mydomains/
N-terminal secondary structure: http://pfam.xfam.org/structure/1WLF
AAA ATPase secondary structure: http://pfam.xfam.org/structure/1D2N