This web page was produced as an assignment for Genetics 564 at UW-Madison in Spring 2014.
What are protein modifications?
Protein modifications are compounds that are added to a protein during and/or after translation. These modifications can affect the overall function of the protein, its localization, and its folding, along with others. Studying these can give insight into the molecular function of a protein and how to better control it. One such modification, phosphorylation, is one of the most important and best characterized. Phosphorylation events are essential for various protein functions and cellular processes [1].
Phosphorylation of PEX1
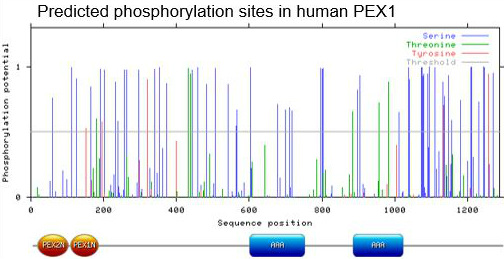
NetPhos 2.0 is a bioinformatic tool that predicts the likelihood of phosphorylation at each phosphorylatable amino acid in a protein sequence: serine, threonine, and tyrosine. This tool assigns a score between zero and one; the larger the number, the more likely it is that this site is phosphorylated in vivo. There are many predicted phosphorylation sites on the PEX1 protein [2]:
Any of these predicted sites could be crucial for proper PEX1 folding and/or function, but none of this data generated is definitive.
Discussion
There are many phosphorylatable sites on PEX1 that have a high phosphorylation potential. This suggests that there are, in fact, phosphorylation events that are crucial for proper function. To investigate this possibility, it would be helpful to determine which of these phosphorylation sites are well-conserved. Those that are well-conserved are probably important for proper function.
[1] "Overview of Post-Translational Modifications (PTMs)." Thermo Scientific. 2014. Web. 12 May 2014. <http://www.piercenet.com/method/overview-post-translational-modification>
[2] Sequence- and structure-based prediction of eukaryotic protein phosphorylation sites. Blom, N., Gammeltoft, S., and Brunak, S. Journal of Molecular Biology: 294(5): 1351-1362, 1999.
Image References
phosphorylation sites generated with: http://www.cbs.dtu.dk/services/NetPhos/
[2] Sequence- and structure-based prediction of eukaryotic protein phosphorylation sites. Blom, N., Gammeltoft, S., and Brunak, S. Journal of Molecular Biology: 294(5): 1351-1362, 1999.
Image References
phosphorylation sites generated with: http://www.cbs.dtu.dk/services/NetPhos/